|
If so, select that channel and explore different parameters and thresholds until the detection looks acceptable.Īlong with the cell detection, QuPath automatically measures all channels in different cell compartments.īecause these measurements are based on the channel names, it is important to have these names established first. The key requirement is that a single channel can be used to detect all nuclei. QuPath’s default Cell detection command can be applied for fluorescence and multiplexed images, not only brightfield. You can either right-click this list or select the ⋮ button and choose Populate from image channels to quickly set these. The classifications currently available are shown under the Annotations tab. We now want to make the channel names available as classifications. 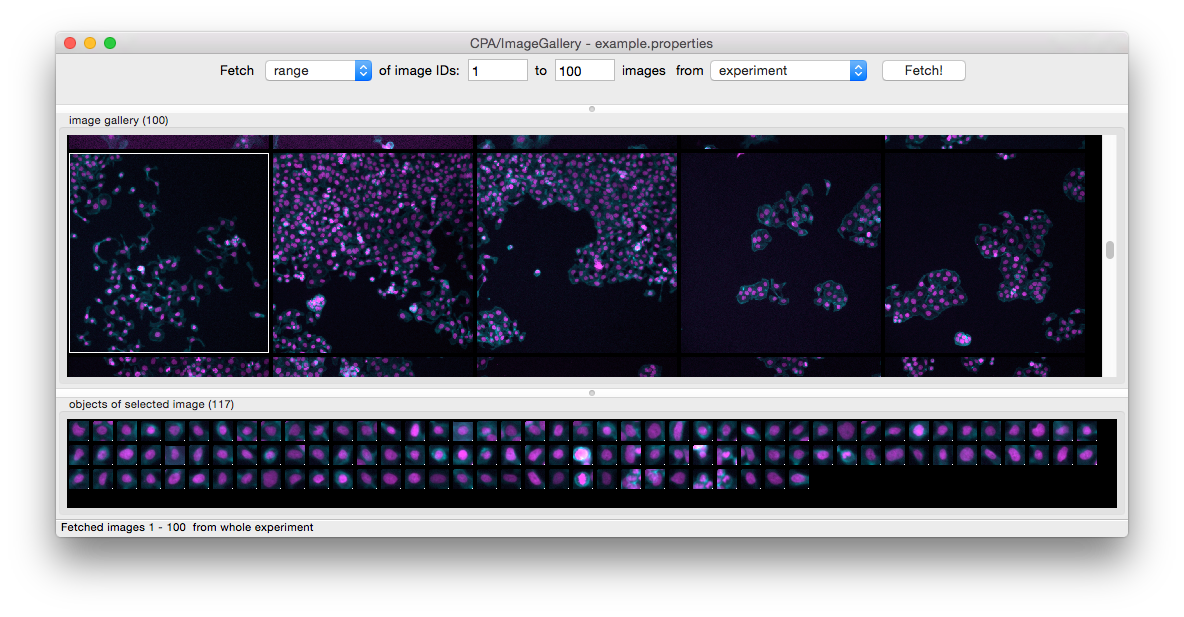
This allows you to reset all the image metadata to whatever was read originally from the file, including the channel names. You can retrieve them later by going to the Image tab, and double-clicking the row that states Metadata changed: Yes. The names can be seen in the Brightness/Contrast dialog, and edited by double-clicking on any entry to change the channel properties. Therefore we usually want them to be short accurate and stripped of any extra text we do not really need. They will also be reused within the names for the cell classifications. 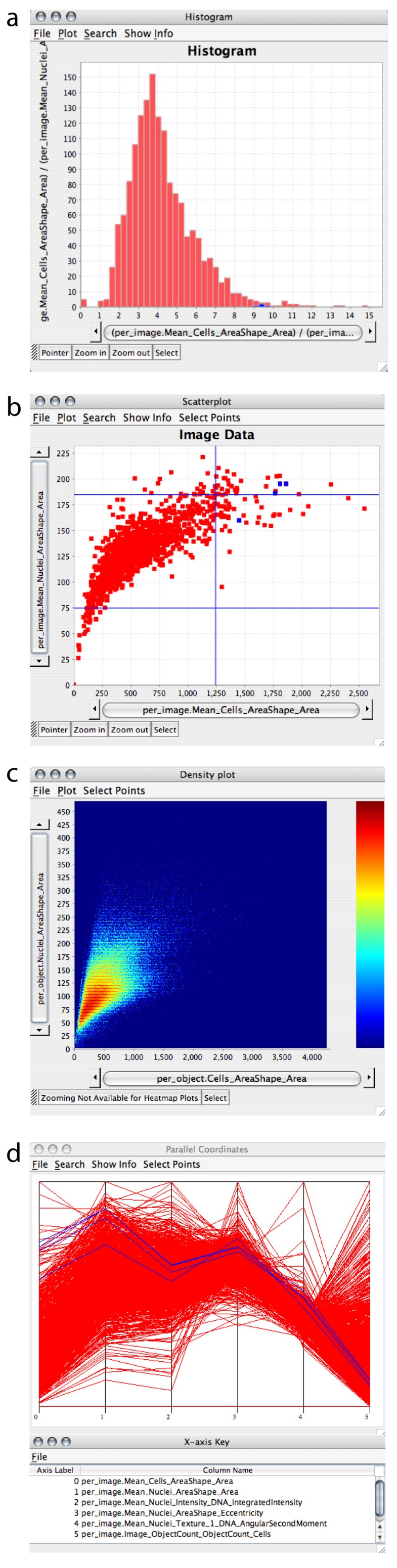
The channel names are particularly important for multiplexed analysis, since these typically correspond to the markers of interest. New and improved methods of segmenting cells in QuPath are being actively explored… Set up the channel names Good cell segmentation is really essential for accurate multiplexed analysis. The Fluorescence type here tells QuPath that ‘high pixel values mean more of something’.Ĭhoosing Brightfield conveys the opposite message, which would cause problems because cell detection would then switch to looking for dark nuclei on a light background. The main thing is to choose the closest match. The type Fluorescence can be used even when not exactly true (e.g. In this case, the best choice is Fluorescence. Set the image type Īs usual when working with an image in QuPath, it is important to ensure the Image type is appropriate. Here, it is really necessary so that classifiers generated along the way are saved in the right place to become available later. Many things in QuPath work best if you create a Project. Step-by-step Before we begin… Create a project containing your images Once this has been done, ‘standard’ QuPath commands and scripts can be used to interrogate the data.Ĭombine the classifiers and apply them to cellsīut first we have a few routine things we need to take care of so that things can run smoothly. We will focus on the main task of identifying each cell, and classifying the cells according to whether they are positive or not for different markers. 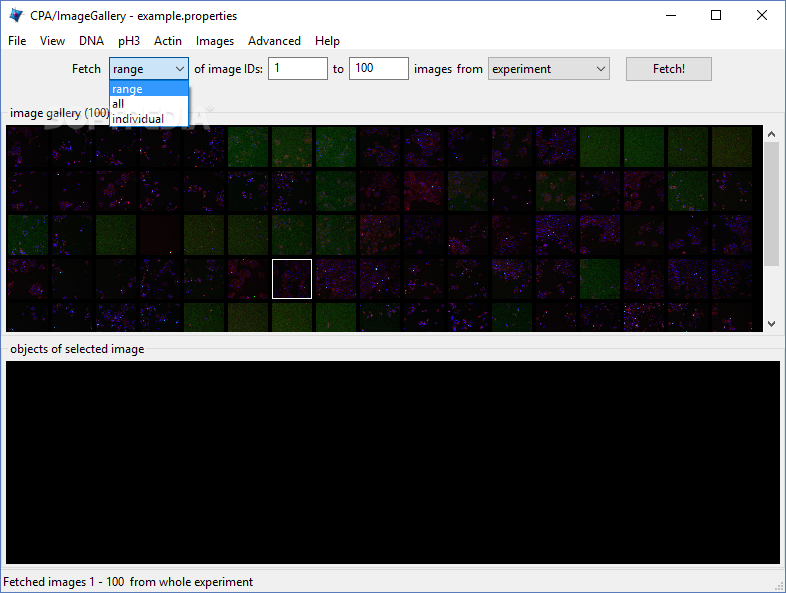
This tutorial outlines the basics of how multiplexed images can be analyzed in QuPath using the sample LuCa-7color_1x1component_data.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
